
While minimal amino acid mutations can yield dramatic enhancements Will benefit from a combination of computational prediction and knowledge Future iterations of protein scaffold discovery and evolution Protein mutants that possess binding functionality over nonfunctional We further explain the necessity for initialĭevelopability by observing a decrease in proteolytic stability of Of Escherichia coli and the ability to evolve bindingįunction upon mutation. Our results support thisĬonnection between developability and evolvability by demonstratingĪ relationship between protein production in the soluble fraction Highly developable (high producibility, stability, solubility) protein Protein engineering endeavors have suggested that starting with a Model analysis suggests a large,ĭisconnected paratope will permit evolved binding function. Providing a 4/6 true positive rate, a 9/11 negative predictive value,Īnd a 4/6 positive predictive value. Regularizationĭeduced which extracted features best predicted binding functionality, Protein scaffolds and reduced into independent components.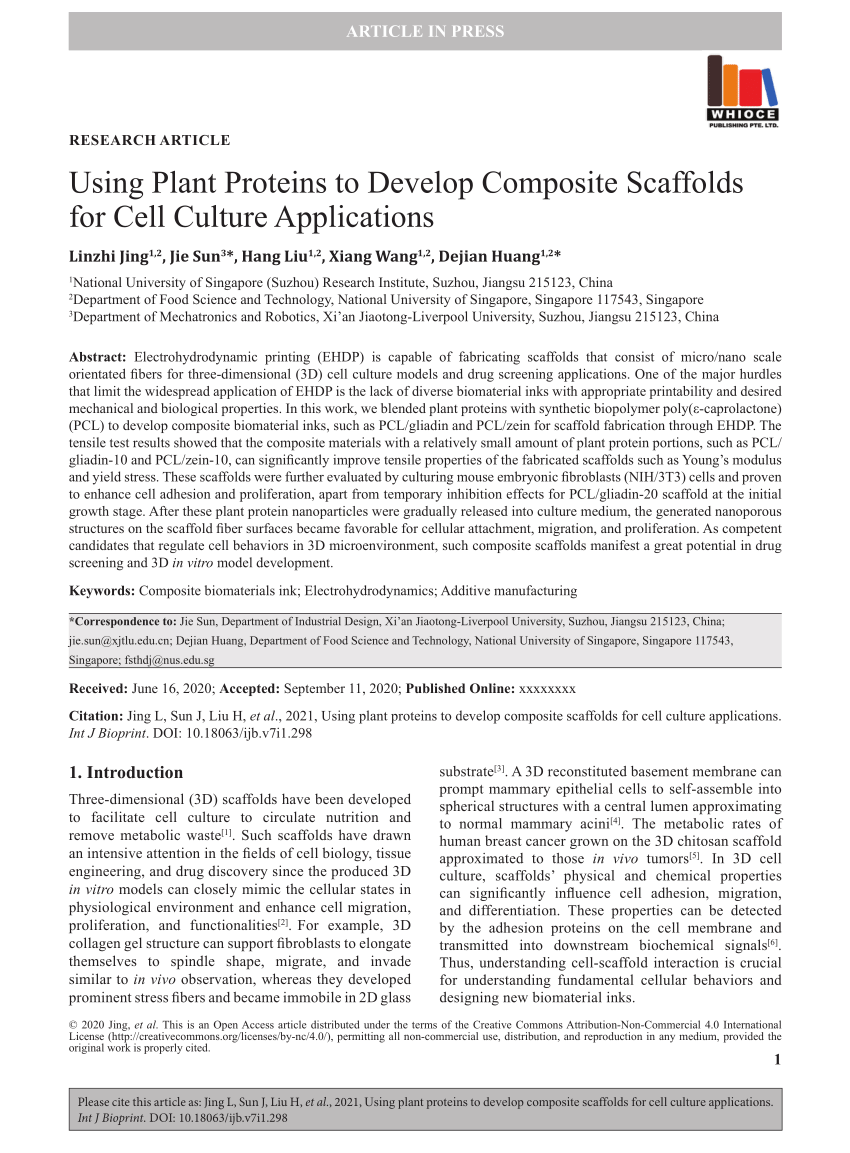 
Topological and biophysical properties were calculated for 787 small Sorting of 10 10 yeast-displayed protein variants. Seven discovery campaigns was determined via magnetic activated cell TheĪbility of 17 small proteins to evolve binding functionality across We present an algorithm to predictĪ protein scaffold’s ability to evolve novel binding functionīased upon computationally calculated biophysical parameters. Scaffolds have been discovered, the underlying properties that permitīinding evolution remain unknown.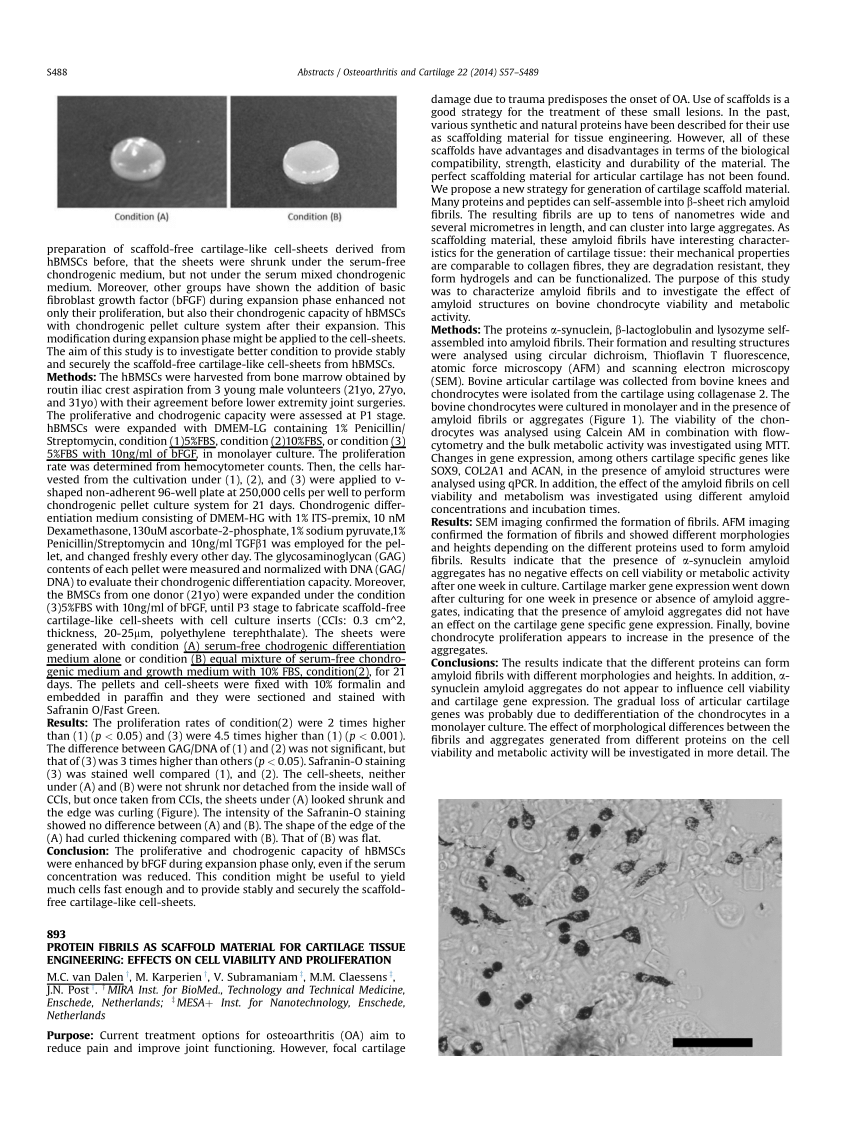 
Protein scaffoldsĪid evolution via a conserved platform on which a modular paratopeĬan be evolved to alter binding specificity. Strategic navigation of a complex mutational landscape. Modalities of interaction of these muteins with proteinases (subtilisin, trypsin and chymotrypsin) were investigated by time course hydrolysis and molecular simulations studies.Molecular recognition function of proteins requires Here, we described the characterization of four muteins containing single/multiple amino acid substitutions at the WSCI reactive site and/or at its proximity. On the other hand, the genetic transformation of cereal plants over-expressing inactive WSCI muteins could represent a possible strategy to improve the nutritional quality of cereal-based foods, without risk of interference with human or animal digestive enzymes. A gene sequence coding for the native WSCI, along with genes coding for muteins with different specficities, could be exploited to obtain transformed non-food use plants with improved insect resistance. The aim of this study was to employ a WSCI molecule as a stable scaffold to obtain recombinant inhibitors with new acquired anti-proteinase activity or, alternatively, inactive WSCI variants. The functional region of WSCI, containing the inhibitor reactive site (Met48-Glu49), corresponds to an extended flexible loop (Val42-Asp53) whose architecture is somehow stabilized by a number of secondary interactions established with a small β-sheet located underneath. In vivo, as suggested for many plant proteinase inhibitors, WSCI seems to play a role of natural defence against attacks of pests and pathogens. In addition to bacterial subtilisins and mammalian chymotrypsins, WSCI inhibits chymotrypsin-like activities isolated from digestive traits of a number of insect larvae. WSCI (Wheat Subtilisin/Chymotrypsin Inhibitor) is a small protein belonging to the Potato inhibitor I family exhibiting a high content of essential amino acid.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
